Quantifying Selection for Pathogenicity and Antibiotic Resistance in Bacteria and Fungi on the ISS – a Microbial Tracking Study – (MT-3C)
Science Objectives
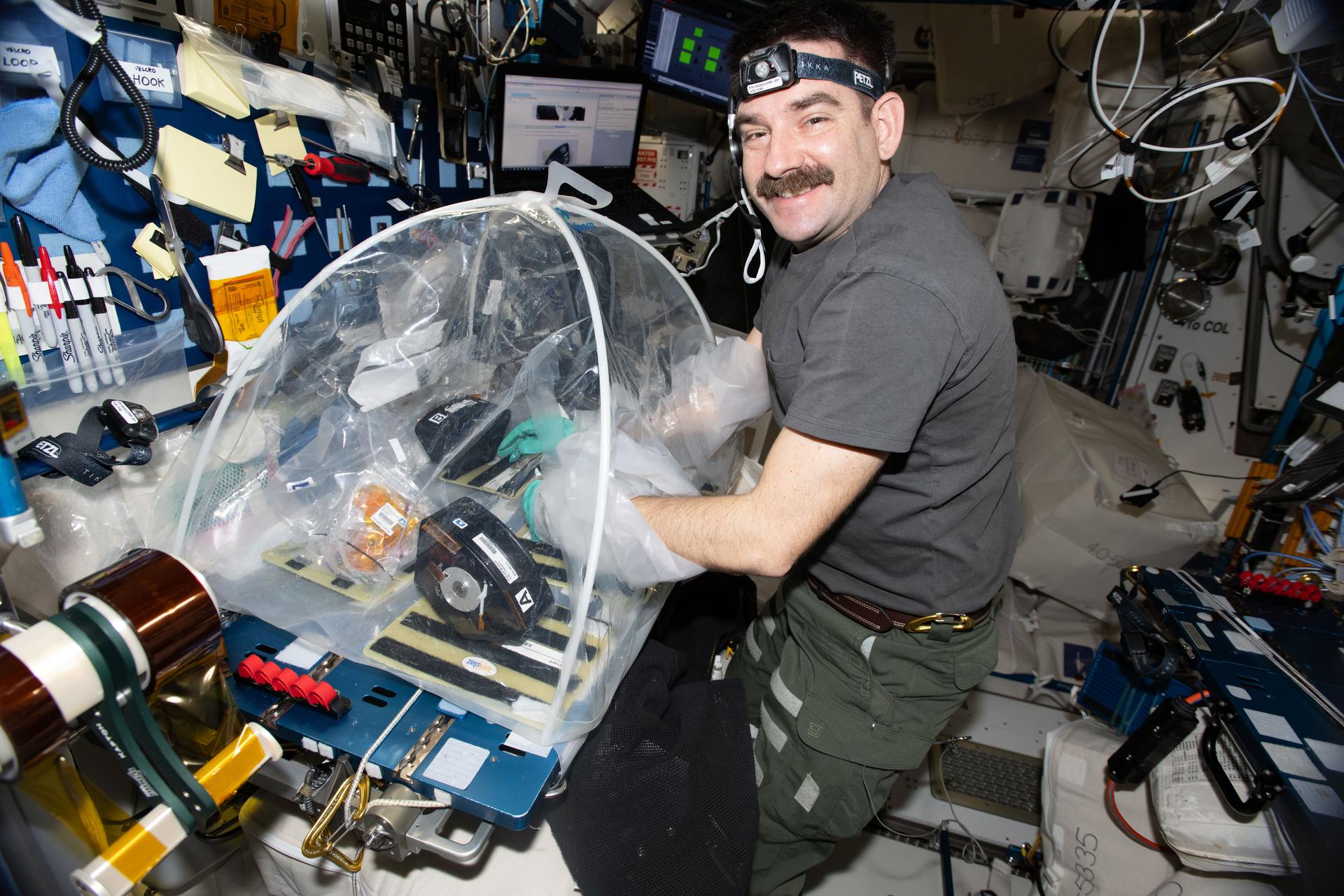
The Quantifying Selection for Pathogenicity and Antibiotic Resistance in Bacteria and Fungi on the ISS – a Microbial Tracking Study (Microbial Tracking-3 or MT-3) investigation continues a series focused on ongoing monitoring of pathogenicity (ability to cause disease) and antibiotic resistance in potentially disease-causing bacteria and fungi present on the International Space Station (ISS). The investigation aims to identify, analyze, and characterize pathogenicity, antibiotic resistance, and genomics to augment the NASA GeneLab with the statistical confidence to characterize microbes associated with closed habitation and predict those that may pose a threat to crew health.
Status
This is the third point in a time series – Microbial Tracking-3 A, B and C. A new batch of sampling kits is going up on SpX-24 to gather more samples for the same study.
Experiment Description
The Quantifying Selection for Pathogenicity and Antibiotic Resistance in Bacteria and Fungi on the ISS (Microbial Tracking-3 or MT-3) investigation hypothesizes that if the International Space Station (ISS) environment promotes microbial stress-survival, there will be an increase in antibiotic resistance and virulence. To test this hypothesis, the research team focuses on the microbiomes of the built environment of spacecraft using existing data (GeneLab) and isolated microbial strains held at JPL/NASA.
Space Applications
This investigation improves understanding of microbes on the space station, including their ability to cause disease and their resistance to antibiotics. It could contribute to developing practical countermeasures to mitigate risk to human health and environmental systems on future space missions. The types of microbes that pose a threat to crew health could be targeted for development of antimicrobial treatment strategies.
Earth Applications
Antibiotic resistance has become a world-wide health concern on Earth. The investigation proposes performing whole genome sequencing on all microorganisms on the space station, which could yield important discoveries related to antibiotic resistance and disease-causing ability. Data also may help improve understanding of how bacteria adapt to built environments, how this adaptation increases disease risk, and what steps could mitigate that risk. An integrated MT-3 database could further enhance various strategies of screening for and identifying specific subsets of microorganisms such as viral and microbial pathogens and those resistant to antibiotics.